Core promoters elements in Xenopus tropicalis
Transcription start sites
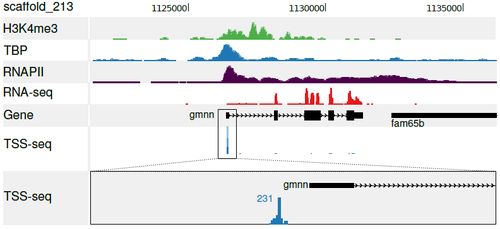
A collection of transcription start sites (TSSs) was obtained by TSS-seq (a CAGE-related approach adapted for deep sequencing) of oocyte and gastrula (stage 10-12) RNA. The TSSs were compared to likely promoter locations using ChIP-sequencing of the general transcription initiation factor TBP and histone H3 K4 trimethylation. The TSSs were used to predict Xenopus core promoter elements, and compare the abundance of these sequence elements in Xenopus and human promoters.
Publications
Van Heeringen, S.J., W. Akhtar, U.G. Jacobi, R.C. Akkers, Y. Suzuki, G.J.C. Veenstra. 2011. Nucleotide composition-linked divergence of vertebrate core promoter architecture. Genome Research, 21, 410-421.
![]() Genome Research Online
Genome Research Online
![]()
Downloads
Visualization and annotation tracks can be used as custom tracks in the UCSC Genome Browser using the links below. These UCSC visualization tracks represent summed read counts at 1bp or 10 bp resolution for TSS-seq and ChIP-seq tracks respectively (available in wiggle format). The Excel table contains the filtered TSS genomic coordinates and the FASTA file (GZip compressed) contains the corresponding promoter sequences from -400 to +100 relative to the TSS.
Assembly: Joint Genome Institute version 4.1 (xenTro2, August 2005).
| |||||||||||||||||
|
|
|||||||||||||||||
The raw data have been deposited in NCBI's Gene Expression Omnibus and are accessible through GEO Series accession number GSE21482
